How it works: CLUSTER
Clustering is a type of analysis that allows us to place similar data objects (users, companies, etc) into groups or clusters. This might be similar behaviours (e.g. power users vs casual users), properties (e.g. demographics), or any other kind of data (e.g. reviews of products, preferred content).
CLUSTER allows users to perform a clustering analysis on any kind of data, making it an extremely powerful method for finding patterns in your data.
But how does it work?!
Automating clustering analysis and explainable AI and the associated machine-learning wizardry 🧙♀️ into one command isn't easy. It requires lots of small innovations and decisions to work seamlessly. 💡
In this page, we will look at what exactly happens when you call CLUSTER. 🤔
TL;DR:
- We use UMAP, a state-of-the-art dimensionality reduction technique, to create a representation of the data that brings similar points closer together and dissimilar points farther apart.
- We use HDBSCAN, a state-of-the-art clustering algorithm, to segment the representation into similar groups or clusters.
- We use PREDICT & EXPLAIN to explain the clusters and understand the patterns in the data.
Overview
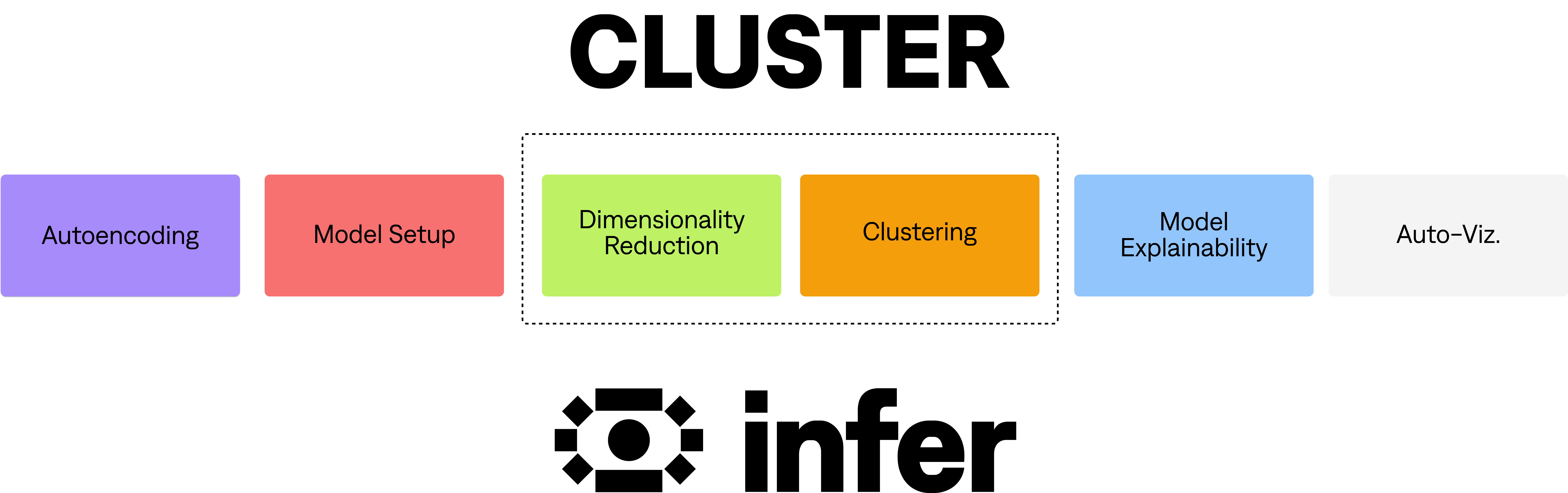
As outlined in the diagram above, there are 6 main processes that occur when calling CLUSTER.
This does not include the added complexity of scaling infrastructure (setting up and running GPU clusters), the parsing of the statement to deconstruct and orchestrate which commands to run, or the interaction with the data sources or data consumers.
This first process of CLUSTER begins when the relevant data has been retrieved from the data source.
These 6 processes for CLUSTER are:
- Autoencoding. Takes the data in its raw form and converts it into a useful format for clustering.
- Model Setup. The configuration of the clustering is decided, based on the autoencoding.
- Dimensionality Reduction. Creates a representation of the data that brings similar points closer together and dissimilar points further apart.
- Clustering. This representation is then segmented into groups of high similarity.
- Model Explainability (EXPLAIN). We use Explainable AI methods to understand what is driving the clustering.
- Auto-viz. Generate relevant insights & visualisations, and return the results to the relevant platform.
In the next sections, we'll outline each process in detail, so you can understand exactly what is happening with confidence.
Autoencoding
Autoencoding is the first step in the CLUSTER process, taking the data and putting it in a suitable form for clustering.
This is largely the same as in PREDICT & EXPLAIN, with a few notable exceptions that will be outlined here.
Categorical Data
- Categorical columns that contain >95% unique values are ignored in the clustering. In contrast,
PREDICTproduces a warning, but will use the column anyway as the overall effect on the model is low. - We do this because the computational memory and time required scales as the number of unique values squared, i.e. adding 10x more unique values results in 100x more time and compute!
- Additionally, including these columns would make the resulting clustering less reliable.
- We one-hot encode the rest of the categorical data.
Numerical Data
- In contrast to
PREDICT, the clustering algorithm is not scale invariant, i.e. making the input values 10x largers may change the outcome. This means we need to rescale all of our numerical features to be comparable. - For clustering numerical data, we first apply robust scaling, to scale the data in a way that is robust to outliers.
- We then apply mix-max scaling, so that the max and min values are between 0 and 1.
- We set missing values to equal 0.0 after robust and min-max scaling, and add a small value to all other values.
Combining
We concatenate the encoded categorical and scaled numerical features into a single representation, which we use as an input into the dimensionality reduction algorithm.
Model Setup
Setting up the model is the second step in the CLUSTER process.
The exact configuration depends on the data and also the user's account type.
Device (GPU/CPU)
CLUSTER works on both GPU and CPU architectures. Using a GPU device allows speedups of 10-100x, making the iterative process of understanding your data even faster.
GPU access is determined on the account level.
Sample & Predict
For datasets larger than 10k, we split the data into two sets - one for training the clustering algorithm, and one for prediction.
The training set is maximum of 10k data points, and the prediction set contains the rest of the data. The idea here is that we can train the model with a high degree of accuracy on 10k data points, while using less memory and compute required for the whole dataset.
After the model is trained, we predict which clusters the rest of the data belongs to. This is much faster than the training process.
Dimensionality Reduction
Dimensionality reduction is the process of reducing the number of variables or features in a dataset while retaining the most relevant information. This can be achieved by transforming high-dimensional data into a lower-dimensional space that still captures the essential patterns and relationships within the data.
One of the main reasons why dimensionality reduction is useful for clustering is that it can help to mitigate the "curse of dimensionality." When working with high-dimensional data, the number of possible feature combinations increases exponentially, making it difficult to identify meaningful patterns and relationships. Dimensionality reduction techniques can help to reduce the complexity of the data, making it easier to cluster similar data points together and identify underlying structures in the data.
Moreover, dimensionality reduction can also help to remove noisy or irrelevant features that may hinder the clustering process. By removing these features, the clustering algorithm can focus on the most important features, resulting in better cluster quality and more interpretable results. Overall, dimensionality reduction is a powerful tool that can improve the effectiveness and efficiency of clustering algorithms, making it an important technique in data analysis and machine learning.
The dimensionality reduction technique used in CLUSTER is called UMAP.
UMAP
UMAP (Uniform Manifold Approximation and Projection) is a machine learning algorithm used for dimensionality reduction. It allows you to take a high-dimensional dataset and represent it in a lower-dimensional space while still preserving the underlying structure and relationships within the data.
To understand how learning lower-dimensional representations of high-dimensional data is possible, consider creation of a video game character. A video game character's appearance is typically created by combining various low-dimensional features such as eye color, nose shape, hair color, and facial structure. These features are combined in different ways to generate a high-dimensional outcome, which is the character's face. UMAP attempts to learn a low-dimensional manifold (eye color, nose) that represents the high-dimensional outcome (faces).
Dimensionality reduction is important because it can help us understand and visualize complex datasets. For example, if you have a dataset with 100 features, it can be difficult to make sense of the relationships between all those features. By reducing the dimensionality of the dataset, we can create a simpler representation that captures the most important information.
UMAP is particularly useful as a preprocessing step before clustering because it can help reveal patterns and structure within the data. Clustering is a technique used to group similar data points together, but it requires that we know what features to look for in order to define those groups. By using UMAP to reduce the dimensionality of the data, we can better visualize the data and potentially identify hidden patterns that we wouldn't have seen otherwise. These patterns can then be used to inform the clustering algorithm and improve its accuracy.
Default Parameters
UMAP has a number of parameters that affect the outcome of the dimensionality reduction. We use the default parameters of UMAP, except for those outlined below.
n_components. We use n_components=5, which represents a reduction to 5 dimensions. We found experimentally that this allows a rich representation of the data without being too computationally expensive.min_dist. We use a minimum distance of 0.0, as recommended for UMAP when the goal is to cluster the representation.metric. We use a cosine similarity metric.n_neighbors. We use themin_cluster_sizeparameter.
We currently don't allow the user to change these parameters directly, but may allow it in the future.
Clustering
Clustering is a data analysis technique that involves grouping similar data points together into clusters based on their features or characteristics. The goal of clustering is to find structure in unlabeled data and gain insights into patterns or relationships that may not be apparent from the raw data.
At Infer, we use a well established and popular algorithm for clustering called HDBSCAN.
HDBSCAN
HDBSCAN stands for "Hierarchical Density-Based Spatial Clustering of Applications with Noise." HDBSCAN is a density-based clustering method that identifies clusters by analyzing the density of data points in the feature space. It works by recursively partitioning the data into subclusters based on density until a cluster is found, or the noise is identified.
HDBSCAN has several advantages over other clustering methods, including the ability to handle clusters of varying shapes and sizes, and the ability to identify noise or outliers. It is particularly useful for datasets with complex structures or high-dimensional data where traditional clustering methods may struggle.
Default Parameters
HDBSCAN has a number of parameters that affect the outcome of the clustering. We use the default parameters of HDBSCAN, except for min_cluster_size, which allows the user to select the minimum size of the clusters (and hence the total number of clusters found).
A user can choose the min_cluster_size like so: CLUSTER(min_cluster_size=100), i.e. this would make the minimum cluster size be of size 100. If the user does not choose, the minimum cluster size is 5% of the dataset size or 10, whichever is larger.
Model Explainability
We run PREDICT and EXPLAIN on the clustering, predicting the cluster ID from the rest of the inputs. This allows us to use all the advantages of PREDICT and EXPLAIN to understand what is driving the clustering.
Auto-viz
The final result is treated identically to PREDICT, producing the same kinds of visualisations as a multi-classification problem.